Blog: News
Whole Genome Sequencing Drives Food Safety
11th May 2017
Whole Genome Sequencing (WGS) for food safety caught my eye recently as it was discussed at the 40th Annual Food Policy Conference in Washington this year. WGS provides the complete DNA code of a specific bacteria. It provides the ability to link a specific strain of Listeria or Salmonella, for example, back to the original source. Quickly linking outbreaks of foodborne illnesses allows investigators to understand how widespread an outbreak is becoming or indeed if more than one primary source is responsible. WGS allows us to link ill patients to likely food sources, by comparing information previously entered in Genome Trakr database, and to bring outbreaks under control more quickly.
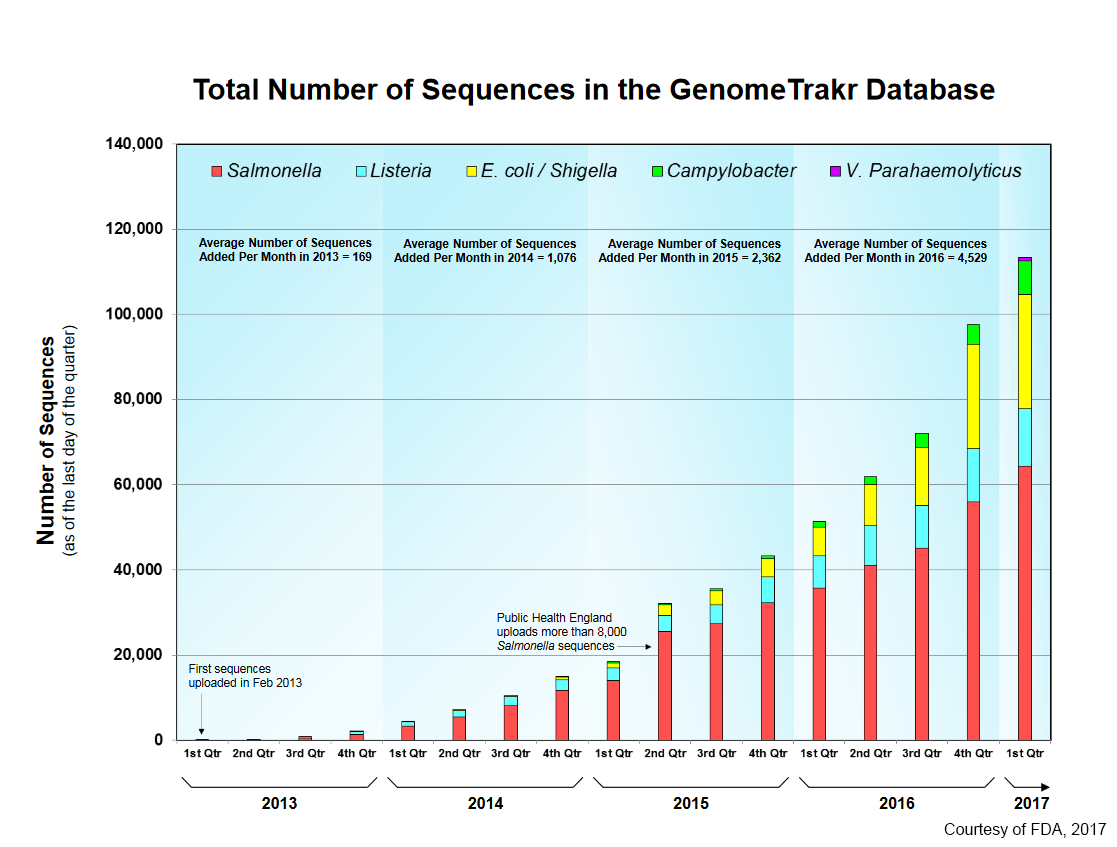
The US FDA has setup a database to track extracted genome sequences for pathogen identification called Genome Trakr. As of May 2017 it had 118,00 sequences on its database and it was growing at a rate of about 4,500 per month.
WGS allows a rapid understanding of foodborne pathogens and provides the potential for us to drive down outbreaks of food borne illnesses by identifying and eliminating the root cause of the contamination. Because WGS is both inexpensive and highly-accurate it is emerging as the next big step forward to further drive good manufacturing practices (GMPs) for food safety.
Further Information on Genome Trkr:
https://www.fda.gov/Food/FoodS...
LIMS within the world of WGS
A LIMS solution can be used to hold sample information and data about that sample. For example the Biomedical Research Centre at Guy's and St Thomas' Hospital in London use Matrix Gemini LIMS the track samples and associated genotyping DNA information extracted from those samples. Whether you are performing whole genome sequencing for the purposes of food safety or extracting DNA to drive studies into personalized medicine managed sample collection and storage facilities using a LIMS, such as these, allow fast recall of the original samples and associated data for further analysis should the need arise.